Molecular Basis of Inheritance: Class 12 Biology NCERT Chapter 6

Key Features of NCERT Material for Class 12 Biology Chapter 6 – Molecular Basis of Inheritance
In the previous chapter: Principles of Inheritance and Variation, You studied inheritance and its laws. In this chapter: Molecular Basis of Inheritance, you will study what is Genetics. You will also get to know more about RNA and DNA.
Prologue to Genetics
How is it conceivable that you resemble your mom and have your dad’s qualities? What makes the kin look and maybe carry on also? Genetics and heredity!
Genetics is characterized from various perspectives. It is characterized as the investigation of qualities, genetic code heredity, and varieties. It is that field of science that portrays how attributes and highlights give from the guardians to their offsprings in each progressive age. The unit of heredity is known as qualities. Before we see more about genetics, we have to comprehend what the genetic materials are.
There are two classes of genetic materials that are answerable for the exchange of data starting with one age then onto the next in creatures:
DNA or deoxyribonucleic corrosive
RNA or ribonucleic corrosive
It is in the DNA or RNA sequences that organic data is put away and passed on.
Quick revision notes
DNA (Deoxyribonucleic Acid) and RNA (Ribonucleic Acid) are two sorts of nucleic corrosive found in living beings. DNA goes about as genetic material in the majority of living beings. RNA likewise goes about as genetic material in certain creatures as in some infections and goes about as courier. It capacities as a connector, auxiliary, and now and again as a reactant particle
The DNA – it is a long polymer of deoxyribonucleotides. A couple of nucleotides is otherwise called base sets. Length of DNA is typically characterized as the number of nucleotides present in it. Escherichia coli have 4.6 x 106 bp and haploid substance of human DNA is 3.3 × 109 bp.
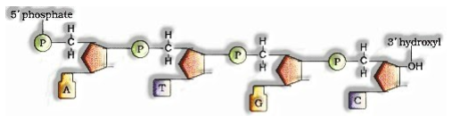
Structure of Polynucleotide Chain
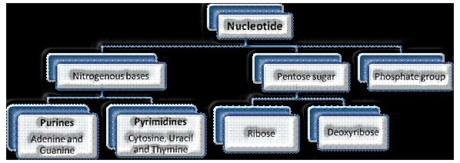
A nucleotide has three components – a nitrogenous base, a pentose sugar (ribose if there should arise an occurrence of RNA, and deoxyribose for DNA), and a phosphate gathering. There are two kinds of nitrogenous bases – Purines (Adenine and Guanine), and Pyrimidines (Cytosine, Uracil and Thymine)
Cytosine is regular for both DNA and RNA and Thymine is available in DNA. Uracil is available in RNA at the spot of Thymine.
A polynucleotide chain
A nitrogenous base is connected to pentose sugar with N-glycosidic linkage to shape to frame a nucleoside. At the point when phosphate bunch is connected 5′- OH of a nucleoside through phosphodiester linkage nucleotide is framed. Two nucleotides are connected through 3′- 5′ phosphodiester linkage to frame dinucleotide. More nucleotide consolidates to shape polynucleotide.
In RNA, nucleotide buildup has extra – OH bunch present at 2′- position in ribose and uracil is found at the spot of Thymine.
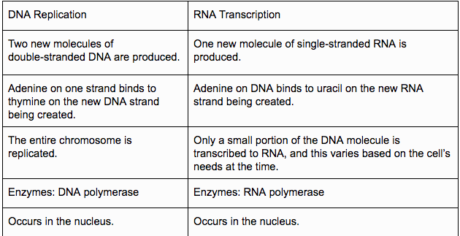
Structure contrasts
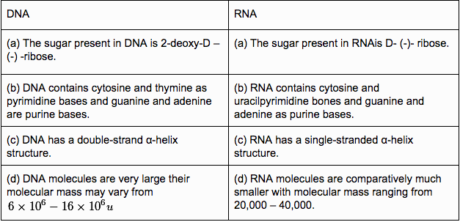
Useful contrasts
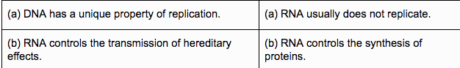
Twofold Helix Model for Structure of DNA-James Watson and Francis Crick, in view of X-beam diffraction information created by Wilkin and Rosalind, proposed this model of DNA.
The quiet highlights of this model are-
a) DNA is made of two polynucleotide chains in which backbone is comprised of sugar-phosphate and bases anticipated inside it.
b) Two chains have hostile to resemble extremity. One 5’à3′ and with 3’à5′.
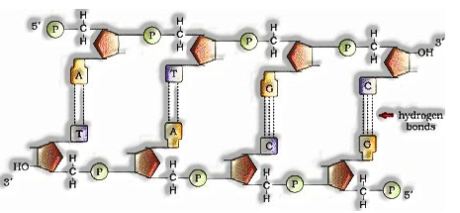
c) The bases in two strands are matched through H-bonds. Adenine and Thymine shapes twofold hydrogen bond and Guanine and Cytosine structures triple hydrogen bonds.
d) Two chains are curled in right gave style. The pitch of the helix is 3.4 nm and about 10 bp in each turn.
e) The plane of one base pair stacks over the other in a twofold helix to give dependability.
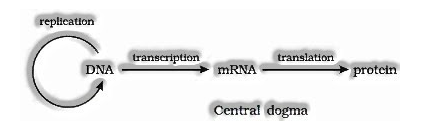
Francis Crick proposed the Central doctrine in sub-atomic science, which expresses that the genetic data streams from DNA — – > RNA — > Protein.
Pressing of DNA helix-
 In prokaryotes, all around the characterized core is missing and adversely accused DNA is consolidated of some decidedly charged proteins called nucleoids.
In prokaryotes, all around the characterized core is missing and adversely accused DNA is consolidated of some decidedly charged proteins called nucleoids.
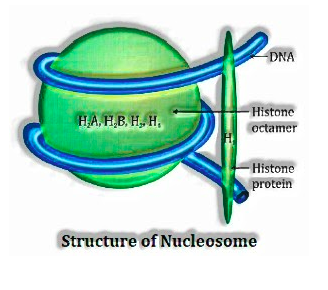
In eukaryotes, histones, decidedly charged protein sorted out to frame 8 atoms unit called histone octamer. Contrarily charged DNA is folded over the histone octamer to shape nucleosome. . Histones are wealthy in the fundamental amino corrosive deposits lysines and arginines. Both the amino corrosive deposits convey positive charges in their side chains.
Single nucleosome contains around 200 base sets. Chromatin is the rehashing unit of the nucleosome.
In core, some area of chromatin are approximately stuffed (and recolours light) and are alluded to as euchromatin. The chromatin that is all the more thickly stuffed and recolours dull are called as Heterochromatin. Euchromatin is transcriptionally dynamic chromatin, while heterochromatin is idle.
The quest for Genetic Material
Changing rule – Frederick Griffith in 1928 led investigate microbes Streptococcus pneumoniae (bacterium liable for pneumonia). There are two kinds of the strain of this microbes, some produce smooth glossy settlements (S) and others produce harsh colonies(R). Mice contaminated with the S strain (destructive) bite the dust from pneumonia disease yet mice tainted with the R strain don’t create pneumonia.
S strain → Inject into mice → Mice kick the bucket
R strain → Inject into mice → Mice live
S strain (heat-executed) → Inject into mice → Mice live
S strain (heat-executed) + R strain (live) → Inject into mice → Mice pass on
Griffith inferred that R strain microscopic organisms have by one way or another changed by heat murdered S strain microorganisms. Some changing standards moved from S strain to R strain and empowered the R strain to integrate a smooth polysaccharide cover and become harmful. This must be because of the exchange of the genetic material.
Biochemical Characterisation of Transforming Principle
Oswald Avery, Colin MacLeod and Maclyn McCarty worked out to decide the biochemical idea of changing standard of Griffith.
They purged biochemicals (proteins, DNA, RNA, and so forth.) from the warmth executed S cells to see which ones could change live R cells into S cells. They found that DNA alone from S microbes made R microscopic organisms become transformed. So, they inferred that DNA is the genetic material.
Trial confirmation that DNA is the genetic material
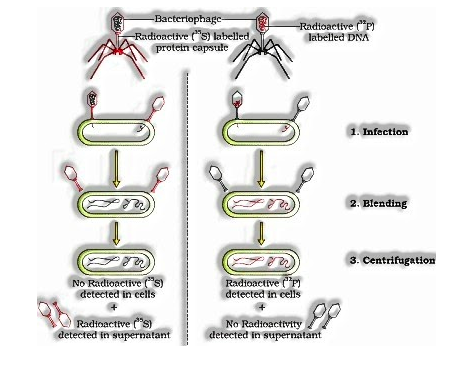
Alfred Hershey and Martha Chases (1952) worked with an infection that contaminates microorganisms called bacteriophages.
In one readiness, the protein part was made radioactive and in the other, nucleic corrosive (DNA) was made radioactive. These two phage arrangements were permitted to taint the way of life of E.coli. Not long after disease, before lysis of cells, the E.coli cells were tenderly unsettled in a blender, to relax the following phage particles and the way of life was centrifuged.
The heavier tainted bacterial cells pelleted to the base and the lighter viral particles were available in the supernatant. It was discovered that when bacteriophage containing radioactive DNA was utilized to contaminate E.coli, the pellet contained radioactivity.
On the off chance that bacteriophage containing radioactive protein coat was utilized to contaminate E.coli, the supernatant contained the vast majority of the radioactivity.
His investigation shows that protein doesn’t enter the bacterial cell and just DNA is the genetic material.
Properties of Genetic Material:
a) It ought to have the option to produce its imitation (replication)
b) It ought to artificially and basically be steady.
c) It ought to give the degree to slow changes (transformation) that are required for advancement.
d) It ought to have the option to communicate as ‘Mendelian Characters’.
• DNA is artificially less receptive however structurally more steady as a contrast with RNA. In this way, DNA is a better genetic material.
• RNA utilized as genetic material just as an impetus and more receptive so less steady. Consequently, DNA has developed from RNA.
Replication of DNA
Watson and Crick proposed that two strands of DNA separate from one another and go about as format for amalgamation of new reciprocal strands. After the culmination of replication every DNA particle would have one parental and one recently orchestrated strand, this technique is called semiconservative replication.
• Messelson and Stahl’s shows test proof of semiconservative replication by developing E .coli on supplement media containing nitrogen salts (15NH4Cl) named with radioactive 15N.
15N was joined into both the strands of DNA and such a DNA was heavier than the DNA gotten from E.coli developed on a medium containing 14N. At that point, they moved the E.coli cells on to a medium containing 14N.
After one age, when one bacterial cell has duplicated into two, they disengaged the DNA and assessed its thickness. Its thickness was middle of the road between that of the heavier 15N-DNA and the lighter 14N-DNA.
This is on the grounds that during replication, new DNA particle with one 15N-old strand and a reciprocal 14N-new strand was shaped (semi-traditionalist replication) thus its thickness is middle of the road between the two.
Replication:
Replication of DNA requires Enzyme DNA polymerase that catalyzes the polymerisation in one strand 5’à3′ simply in the wake of loosening up with the assistance of Helicase catalyst. In this way, replication in one stand is nonstop and another strand it is broken to incorporate Okazaki parts that are consolidated by compound DNA ligase.
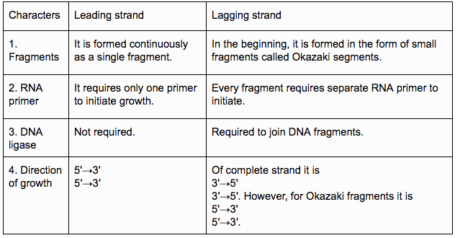
Contrasts among driving and Lagging strands for DNA replication.
Record
It is the way toward duplicating genetic data from one strand of DNA into RNA. In record, just one portion of DNA and just one strand is replicated in RNA. The Adenosine structures base pair with Uracil rather than Thymine.
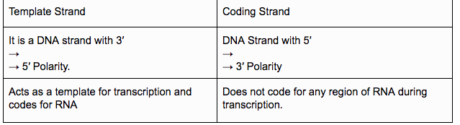
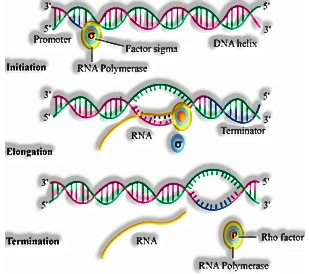
- Transcription of DNA incorporates an advertiser, the auxiliary quality and an eliminator. The strands that have extremity 3’à5 go about as format and called layout strand and another strand is called a coding strand.
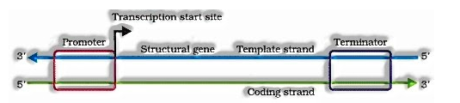
 The advertiser is situated at 5′ end and that dilemma the protein RNA polymerase to begin the record. Sigma factor likewise helps in initiation of record. The eliminator is situated at 3’end of coding strand and as a rule, characterizes the finish of record where rho factor will tie to end the record.
The advertiser is situated at 5′ end and that dilemma the protein RNA polymerase to begin the record. Sigma factor likewise helps in initiation of record. The eliminator is situated at 3’end of coding strand and as a rule, characterizes the finish of record where rho factor will tie to end the record.
Exons are those sequences that show up in develop and handled RNA. Exons are hindered by introns. Introns don’t show up in develop and prepared RNA.
In eukaryotes, there are three distinctive RNA polymerase chemicals I, II and III, they catalyze the blend of a wide range of RNA.
RNA polymerase I – rRNAs
The RNA polymerase II – mRNA
RNA polymerase III – tRNA
The m-RNA give the format, tRNA brings the amino acids and read the genetic code, the rRNA assume auxiliary and synergist job during interpretation
The essential record contains both exon and intron and is non-practical. It experiences the way toward grafting where introns are expelled and exons participate in a characterized request.
The hnRNA (heterogeneous atomic RNA) experience extra preparing called as a topping and following. In topping in uncommon nucleotide (methylguanosine triphosphate) to the 5’end of hnRNA. In following polyadenylatelate tail is included at 3’end in a format at autonomous way.
Genetic Code:
Genetic Code is the relationship of amino acids sequence in a polypeptide and nucleotide/base sequence in mRNA. It coordinates the sequence of amino acids during the union of proteins.
George Gamow proposed that genetic code ought to be a blend of 3 nucleotides to code 20 amino acids.
H.G. Khorana created substance strategy to blending RNA particles with a characterized mix of bases.
Marshall Nirenberg’s without cell framework for protein amalgamation, at last, helped the code to be unravelled.
Striking highlights of Genetic Code are
- The code is a triplet. 61 codons code for amino acids and 3 codons don’t code for any amino acids called stop codon (UAG, UGA and UAA).
- A codon is unambiguous and explicit, code for one amino corrosive.
- The code is degenerate. Some amino acids are coded by more than one codon.
- The codon is touchingly perused in mRNA with no accentuation.
- The codon is almost widespread. AUG has double capacities. It codes for methionine and goes about as initiator codon.
Transformations and Genetic code
A difference in single base pair (point transformation) in the sixth situation of Beta globin chain of Hemoglobin results because of the difference in amino corrosive buildup glutamate to valine. These outcomes into an unhealthy condition called sickle cell pallor.
Inclusion and erasure of three or its various bases embed or erase one or different codons thus at least one amino acids and perusing outline stays unaltered starting there onwards. Such changes are called outline move addition or cancellation transformations.
tRNA–the Adapter Molecule
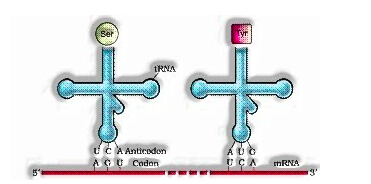
The t-RNA called as connector atoms. It has an anticodon circle that has puts together integral to code present with respect to mRNA and furthermore has an amino corrosive acceptor to which amino corrosive ties. t-RNA is explicit for every amino acids.
The optional structure of t-RNA is portrayed as clover-leaf. In real structure, the t-RNA is a reduced atom which resembles rearranged L.
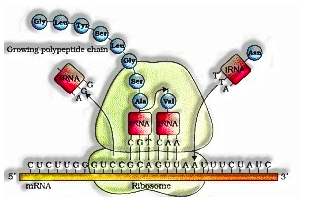
Interpretation process: Translation is the procedure of polymerisation of amino acids to frame a polypeptide. The request and sequence of amino acids are characterized by the sequence of bases in the mRNA. Amino acids are joined by peptide bonds. It included advances
a) Charging of t-RNA.
b) Formation of peptide bonds between two charged tRNA.
- The beginning codon is AUG. An mRNA has some extra sequence that is not interpreted called untranslated locale (UTR).
- For commencement ribosome ties to mRNA toward the beginning codon. Ribosomes move from codon to codon along mRNA for prolongation of the protein chain. Toward the end discharge factors ties to the stop codon, ending the interpretation and arrival of polypeptide structure ribosome.
Guideline of Gene Expression:
All the qualities are not required continually. The qualities required just in some cases are called administrative qualities and are made to work just when required and remain non-utilitarian at different occasions. Such managed qualities, subsequently required to be turned ‘on’ or ‘off’ when a specific capacity is to start or stop.
The Lac Operon
Lac operon comprises of one administrative quality (I ) and three auxiliary qualities (y,z and a). Quality I code for the repressor of the lac operon. The z quality code for beta-galactosidase, that is answerable for hydrolysis of the disaccharide, lactose into monomeric units, galactose and glucose. Quality y code for permease, which expands penetrability of the cell. Quality an encode for transacetylase.
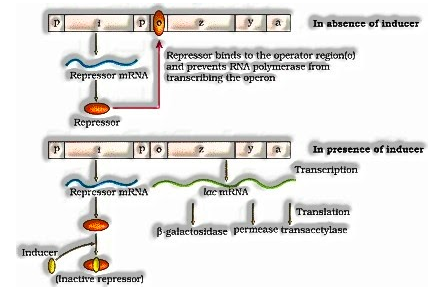
Lactose is the substrate for catalyst beta-galactosidase and it manages to turn on and off of the operon, so it is called inducer.
Guideline of Lac operon by repressor is alluded as negative guideline. Activity of Lac operon is likewise heavily influenced by positive guideline.
Human Genome Project was propelled in 1990 to discover the total DNA sequence of human genome utilizing genetic building procedure and bioinformatics to detach and clone the DNA section for deciding DNA sequence.
Objective of HGP-
a) Identify all the qualities (20,000 to 25,000) in human DNA.
b) Determine the sequence of the 3 billion substance base matches that make up human DNA.
c) Store this data in an information base.
d) Improve devices for information examination.
e) Transfer related data to different divisions.
f) To address the legitimate, moral and social issues that may emerge because of the undertaking.
- The venture was composed by the US Department of Energy and the National Institute of wellbeing.
- The technique included the two significant methodologies initially recognizing all the qualities that express as RNA called Express sequence tags(EST). The second is the sequencing all arrangement of the genome that contained all the coding and non-coding sequence called sequence Annotation.
Remarkable highlights of Human Genome:
a) The human genome contains 3164.7 million nucleotide bases.
b) The normal quality comprises of 3000 bases, yet measures shift extraordinarily, with the biggest realized human quality being dystrophin at 2.4 million bases.
c) Less than 2 per cent of the genome codes for proteins.
d) Repeated sequences make up an exceptionally enormous bit of the human genome.
e) Repetitive sequences are stretches of DNA sequences that are rehashed ordinarily, now and then hundred to multiple times.
f) Chromosome 1 has the most qualities (2968), and the Y has the least (231).
g) Scientists have recognized about 1.4 million areas where single-base DNA contrasts (SNPs – single nucleotide polymorphism) happen in people.
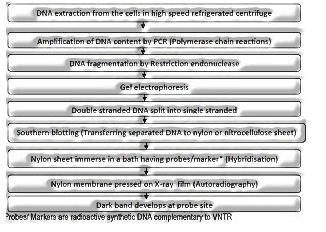
DNA fingerprinting is an exceptionally fast approach to analyze the DNA sequence of any two-person. It incorporates distinguishing contrasts in some particular area in DNA sequence called as tedious DNA in light of the fact that in this district, a little stretch of DNA is rehashed ordinarily.
Contingent on the base organization, length of portion and number of monotonous units satellite DNA is arranged into numerous classes.
Polymorphism in DNA sequence is the reason for genetic planning of human genome just as fingerprinting.
The procedure of fingerprinting was at first evolved by Alec Jeffrey. He utilized a satellite DNA as test to so high polymorphism was called Variable Number of Tendon Repeats (VNTR).
Questions
Q1: What keeps the two strands of DNA connected to one another?
Hydrogen bond
Covalent bond
Ionic bond
The bond between the sugar atom and the phosphate particle
Sol. The right answer is the alternative “a”. The hydrogen bonds between corresponding bases in the two DNA strands keep it together.